Our preprint is out in bioRxiv: https://www.biorxiv.org/content/10.1101/844852v1
Filtering raw genomic datasets is essential to avoid chimeric assemblies and to increase the validity of sequence-based biological inference. BlobToolKit extends the BlobTools1/Blobology2 approach to simplify interactive and reproducible filtering.
BlobToolKit is comprised of four components:
- BlobToolKit Viewer allows browser-based interactive visualisation and filtering of preliminary or published genomic datasets even for highly fragmented assemblies.
- BlobTools2 is a command-line program to convert assemblies and analysis results into datasets that can be further processed using BlobTools2 and/or visualised in the Viewer.
- The BlobToolKit Specification features a formal schema and validator for the JSON-based BlobDir format used by BlobTools2 and the Viewer.
- The BlobToolKit Pipeline is a configurable Snakemake pipeline that automates all steps from retrieving public datasets through running analyses and generating a BlobDir dataset with BlobTools2, ready for visualisation in the Viewer.
The Viewer features multiple views and data export options that dynamically update as filter parameters and selections are modified (Figure 1).
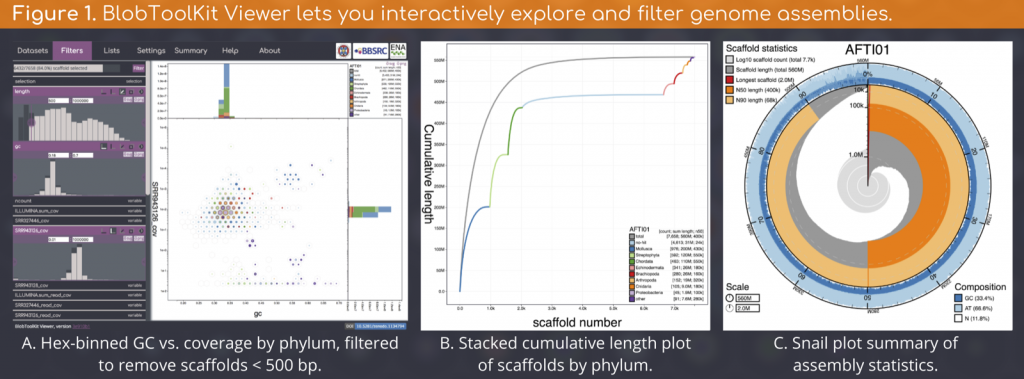
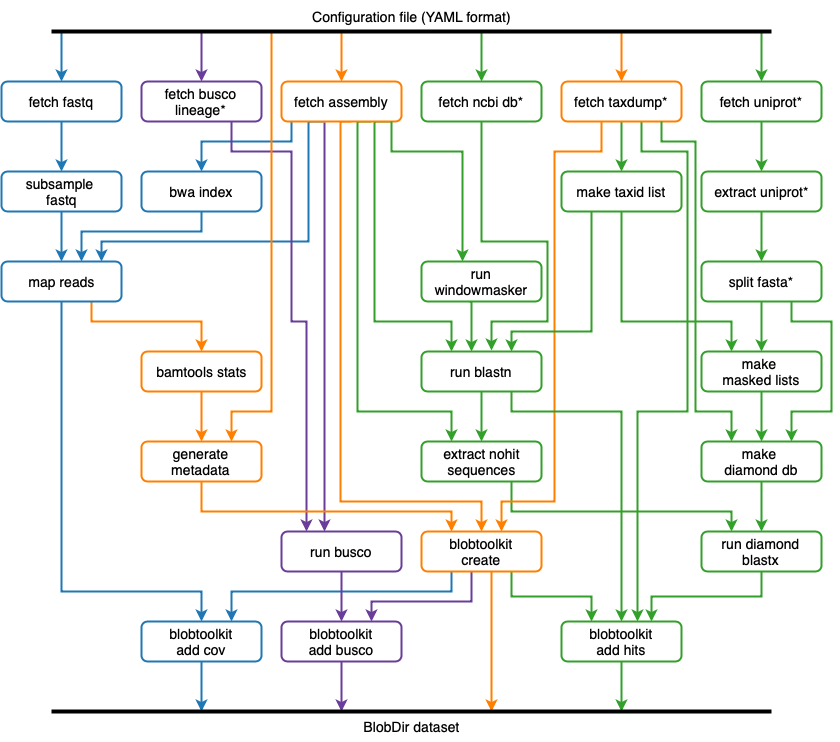
We are running the BlobToolKit pipeline (Figure 2) on all public (INSDC registered) eukaryote genome assemblies and making the results available on a public instance off the Viewer at blobtoolkit.genomehubs.org/view.
1Kumar et al. 2013. Blobology: exploring raw genome data for contaminants, symbionts and parasites using taxon-annotated GC-coverage plots. Frontiers in Genetics, 4:237
2 Laetsch & Blaxter 2017. BlobTools: Interrogation of genome assemblies [version 1; referees: awaiting peer review]. F1000Research, 6:1287 (doi: 10.12688/f1000research.12232.1)

